1.2 INPUTNEWLEAD attempts to connects all fragments in the input file, that carry H-atoms explicitly. The following input formats are recognized automatically: Columbia (Macromodel), MOL2 (Sybyl), and CAR (Accelrys) archive files. The same format is then used for the output file or files [2]. The graphic interface for InsightII creates a suitable input transparently. You can also define a "receptor" - one or more molecules which are not connected to the spacers but define a region inaccessible to them. stand-alone use Write the input file for NEWLEAD from your modelling program, and give it a specific name. The default name is "nld.inp". The receptor is defined by colour. The fragments must be coloured normally and no adjacent atoms should be red. You must define the receptor by colouring all its atoms in red. |
||
|
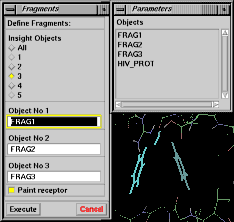
Accelrys interface In InsightII, you can define the fragments and the receptor graphically. Activate the Fragments command of the Custom menu (Fig 1), after the fragments and the receptor (if any) have been defined and are displayed on the screen in the desired positions. As a default, all objects on the screen are considered as fragments, and the button "All" is checked. If a receptor is present, check the button corresponding to the number of fragments. Then select the fragments by clicking on them or by picking their names from the menu. The remaining structures are interpreted as the receptor. Select the "Paint receptor" button to see the receptor turn red. Clicking the "Execute" button completes the operation. |
|